Over the past 5 years, Explicyte has supported several cancer immunotherapy programs aimed at inducing or enhancing a specific anti-tumor immune response. Here’s a quick overview of our capacities for the advancement of novel cancer vaccines (peptide-based, DC-based, DNA- or RNA-based), cytokine therapies, oncolytic virus-based therapies, immune modulators (such as STING agonists), and immune adjuvants.
Specific Tumor Antigen Presentation: In vitro Efficacy & MoA Studies with a dedicated DC-mediated T-cell Killing Assay
We designed a cell-based system to assess the effectiveness of compounds aimed at enhancing the proficiency of antigen-presenting cells (APC), such as dendritic cells (DC), to capture and cross-present the tumor antigens, and the subsequent priming of CD8+ cytotoxic T cells against cancer cells. The assay provides dynamic flexibility, thus fitting different typologies of test compounds, with different modes of action and treatment modalities.
CASE STUDY
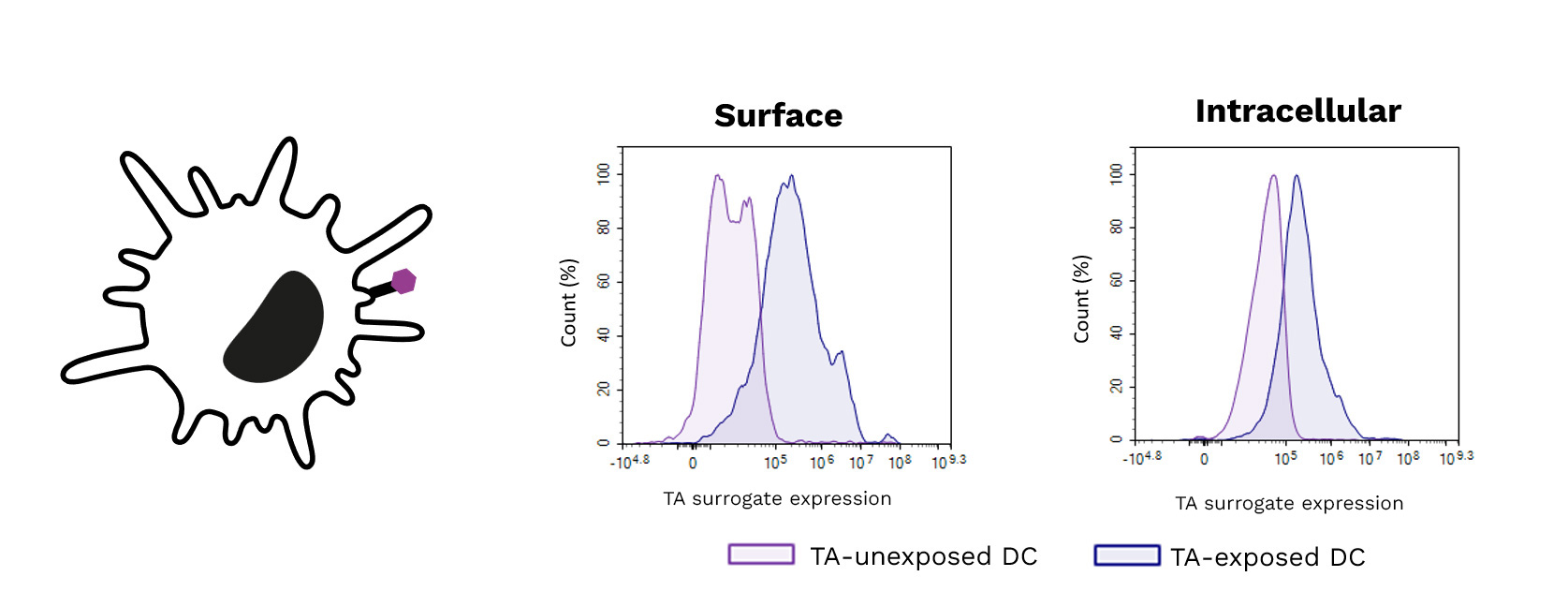
Exposure of DCs to a target tumor antigen (TA) peptide enhances their antigenic presentation shown by the increase of its surrogate expression (surface (left) and intracellular (right)) using specific antibodies.
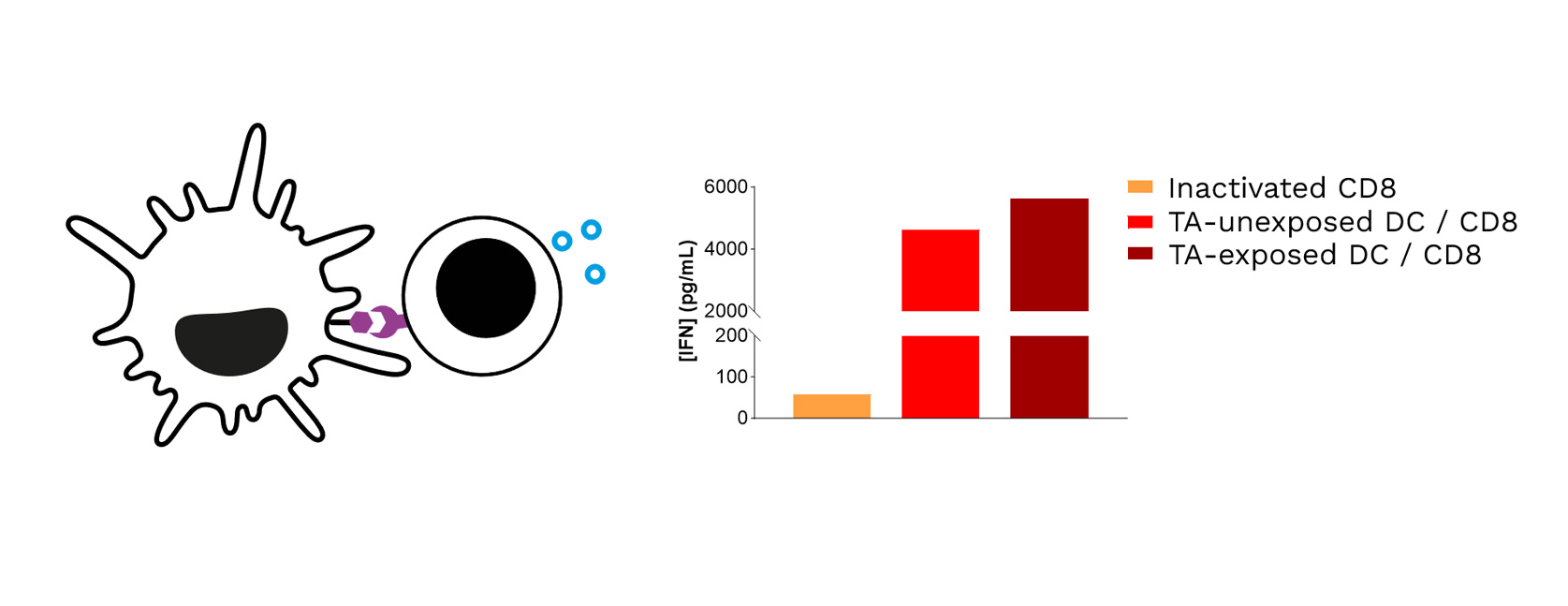
Target TA-exposed DCs lead to optimized priming of CD8 T cells, as shown through the increased IFNγ levels released in CD8 T cell / DC co-cultures.
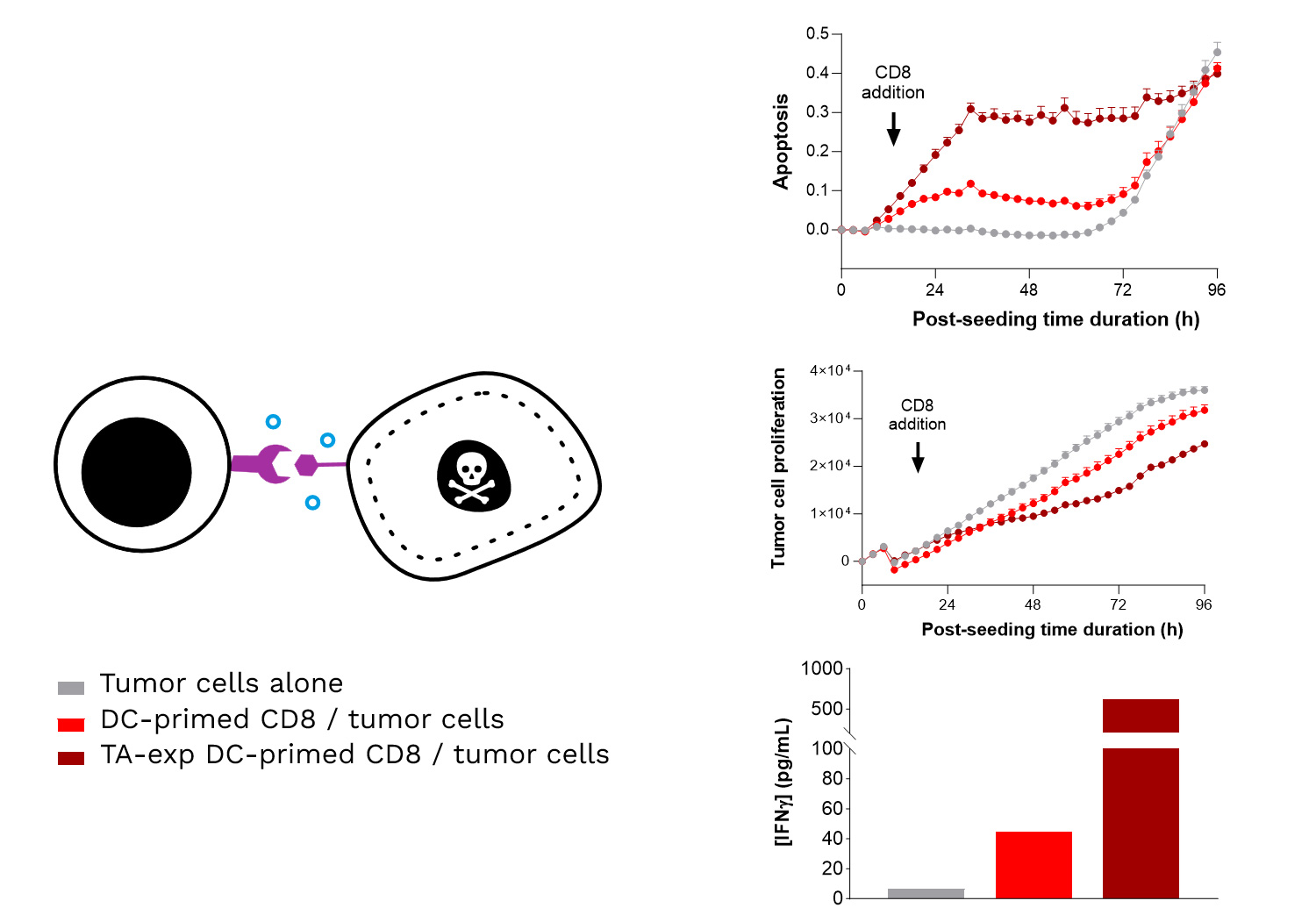
Specific TA-exposed DC-mediated priming of CD8 T cells results in the induction of an adaptive effector T cell-mediated killing towards SK-MEL-5 tumor cells (target TA-positive). Real-time monitoring of SK-MEL-5 tumor cell killing mediated by CD8 T cells, primed by either TA-exposed or unexposed DCs, where apoptosis and tumor cell count were monitored over ~4 days and analyzed. In addition, IFNγ release by CD8 T cells is shown to be increased upon their TA-exposed DC priming, compared to TA-unexposed DCs.
TAKE HOME MESSAGE: Enhancing DC-mediated tumor antigen presentation induces a specific, optimized, adaptive effector T cell-mediated killing response.
How does it work?
A step-by-step overview of our tumor antigen-specific T-cell mediated tumor killing assay:
- Choice of the right tumor cell lines based on their expression of the target antigens and their representativeness of the cancer indications of interest (>100 in-house human tumor cell lines)
- Isolation of PBMC-originating immune populations required (monocytes, CD8) for autologous co-cultures
- Generation of monocyte-derived APC such as differentiated & mature DCs
- DC – CD8 T cell co-cultures for DC-mediated priming and activation of CD8 cells
- CD8 T cell – target tumor cell co-cultures to capture the tumor-targeted cytotoxic response
- Adequate treatment windows in line with the expected mechanisms of action and eventual promising combination treatments that could hold potential for improving anti-tumor response
- Relevant readouts to capture each cell component (DC cytokines and surface markers, CD8 cytokines and surface markers, target tumor cell apoptosis/proliferation…)
More data ? Some questions ? Contact our team !
Our capacities for the identification & Validation of Target Antigen Expression in Tumor Samples
Starting from FFPE tumor specimens, our translational team can explore and/or validate the expression of target tumor antigens across a set of cancer indications, using various platforms:
- Single-Cell RNA-Seq to identify & quantify tumor antigen expression, and investigate tumor antigen heterogeneity and the immune landscape
- Xenium Single-Cell Transcriptomics to explore tumor antigen patterns and tumor heterogeneity with respect to the immune microenvironment (immune infiltration, response, gene expression profiles and signatures…)
- GeoMx Digital Spatial Profiling for comprehensive antigen mapping within the tumor microenvironment (TME)
- Digital Pathology for precise and multiplexed analysis of antigen expression with the TME – providing a comprehensive view of the tumor antigenic profile and how it could interact with other components of the TME